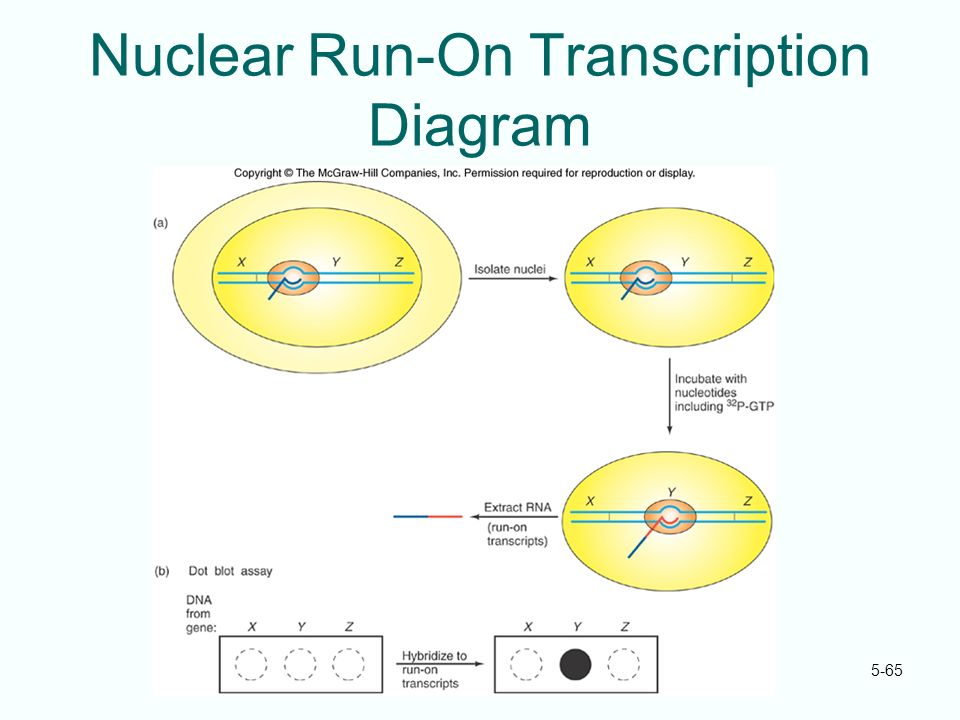
Comparison of the conventional run-off transcription and the ribozyme... | Download Scientific Diagram

Identifying Transcription Error-Enriched Genomic Loci Using Nuclear Run-on Circular-Sequencing Coupled with Background Error Modeling - ScienceDirect
The size of this run-off transcript locates the transcription start site, and the amount of this transcript reflects the efficiency of transcription. -triyambak.org - Triyambak Life Sciences

Identifying Transcription Error-Enriched Genomic Loci Using Nuclear Run-on Circular-Sequencing Coupled with Background Error Modeling - ScienceDirect

Quantification of nascent transcription by bromouridine immunocapture nuclear run-on RT-qPCR | Nature Protocols

New Chromatin Run-On Reaction Enables Global Mapping of Active RNA Polymerase Locations in an Enrichment-free Manner | ACS Chemical Biology
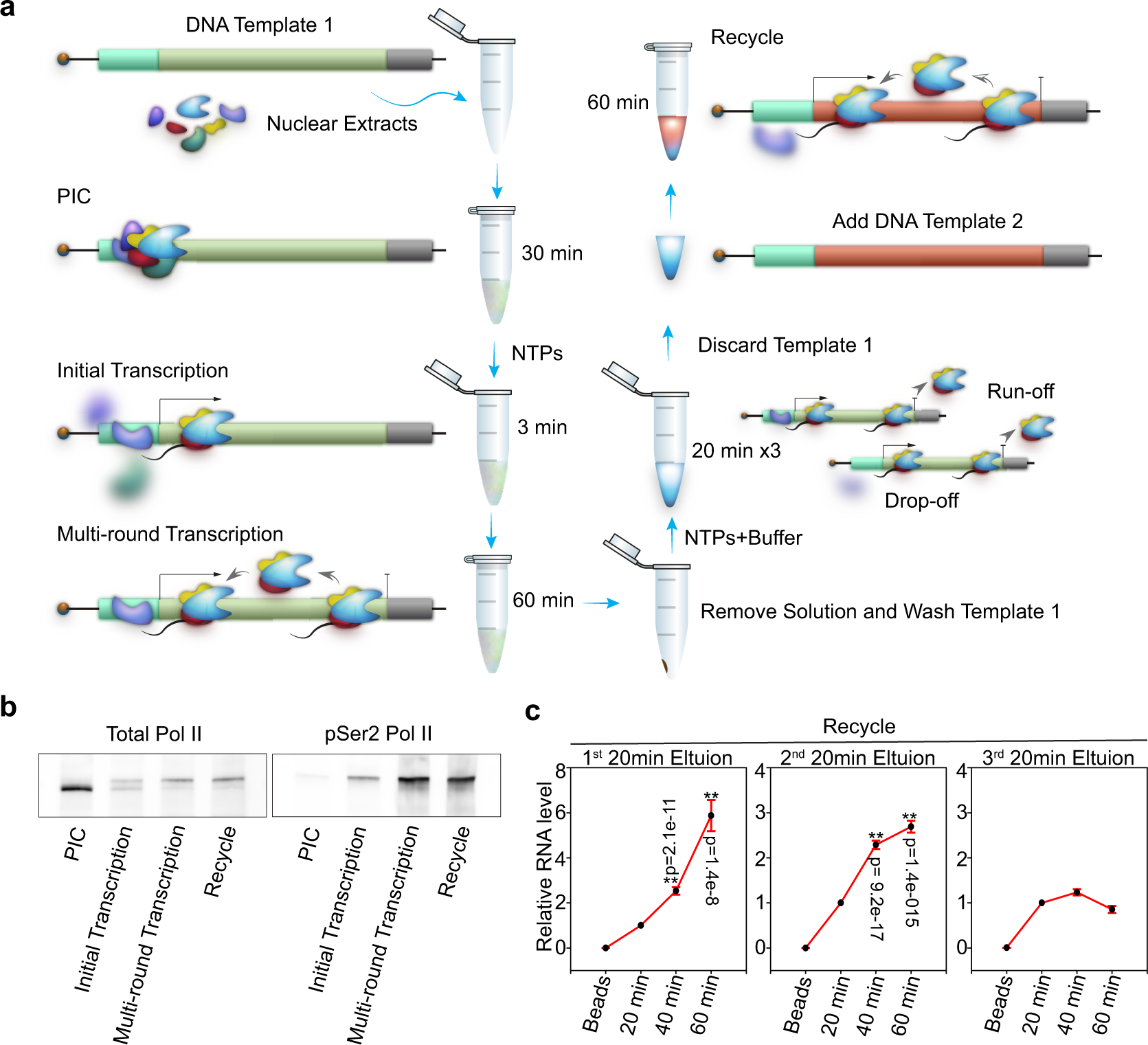
Transcription recycling assays identify PAF1 as a driver for RNA Pol II recycling | Nature Communications
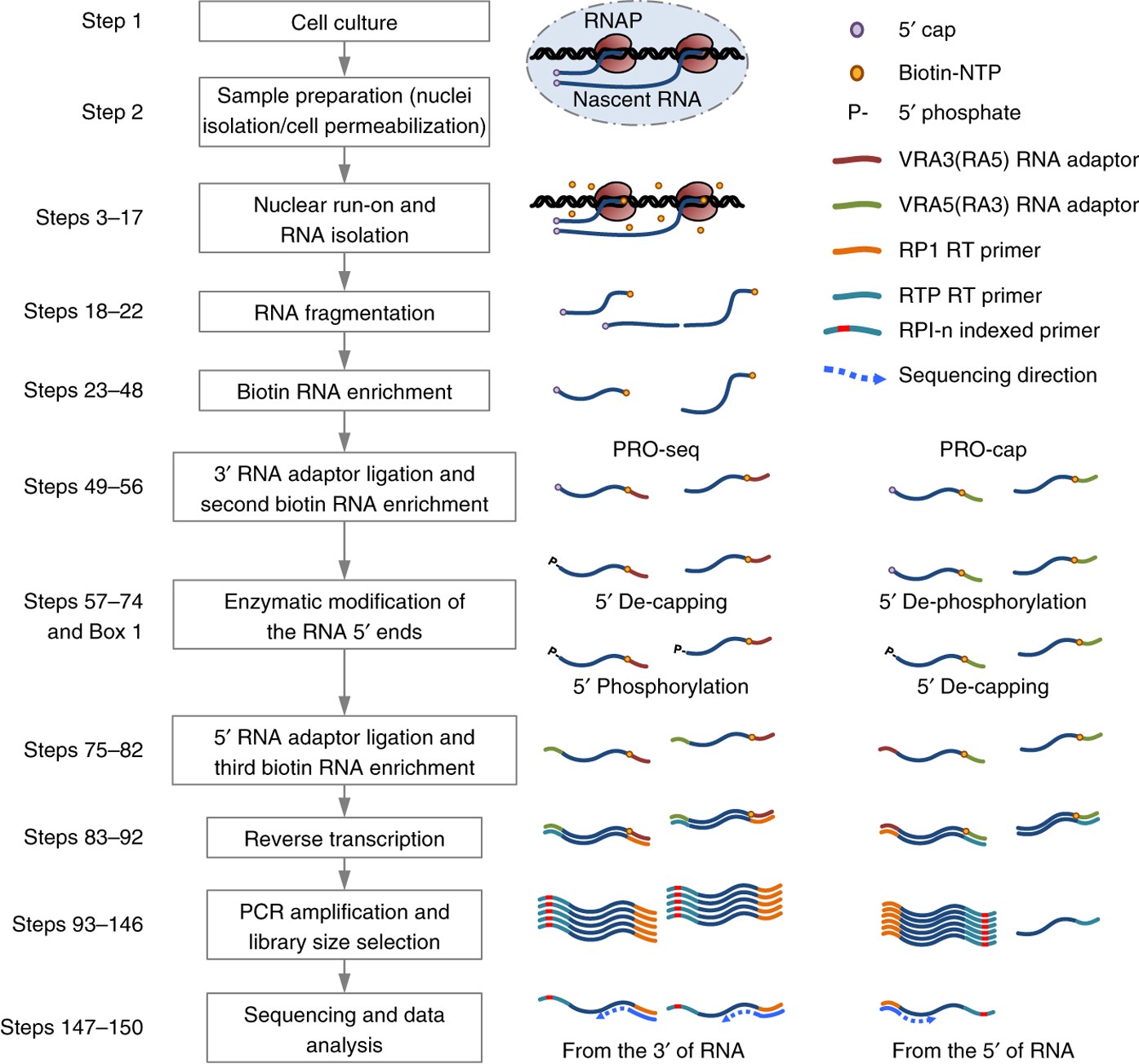


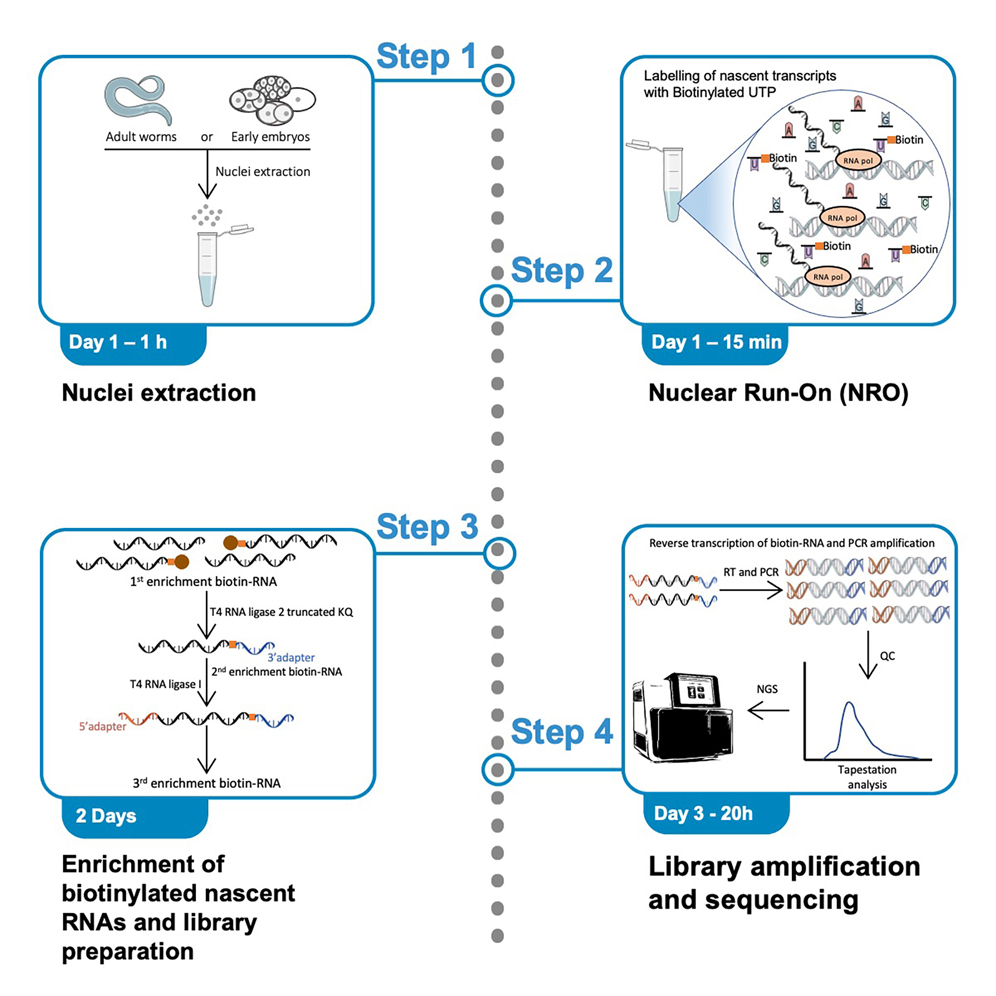
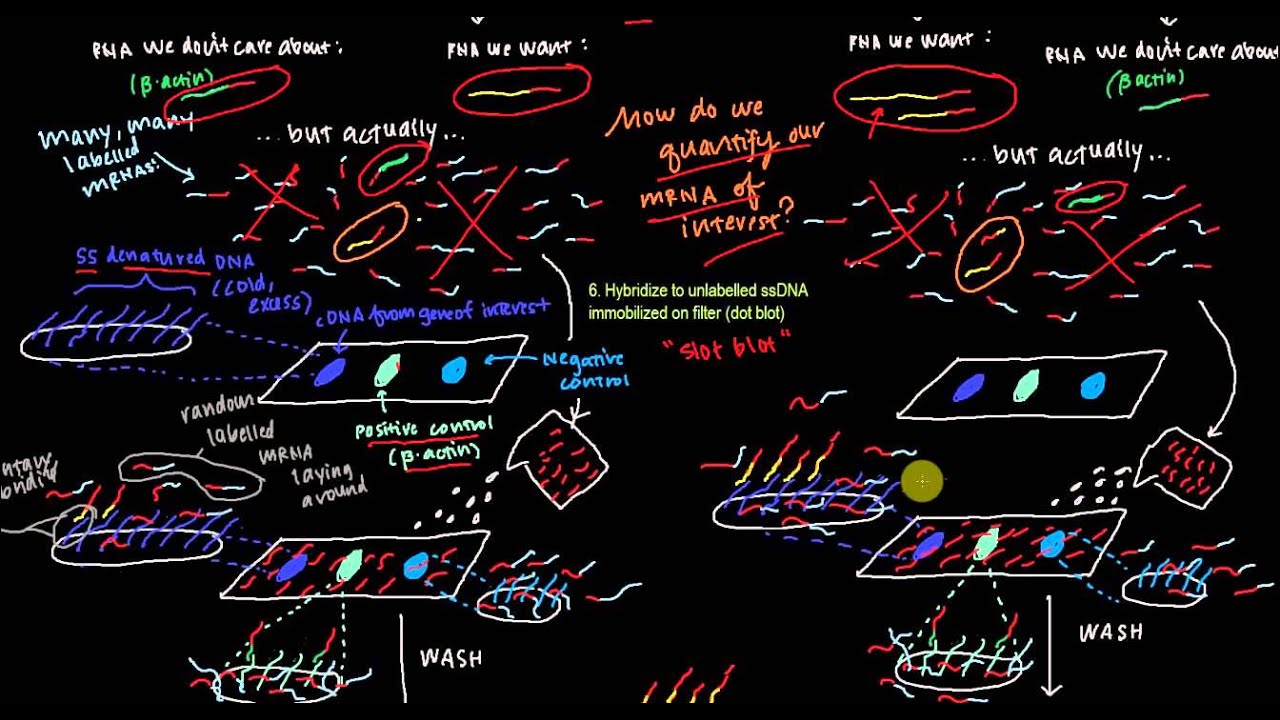
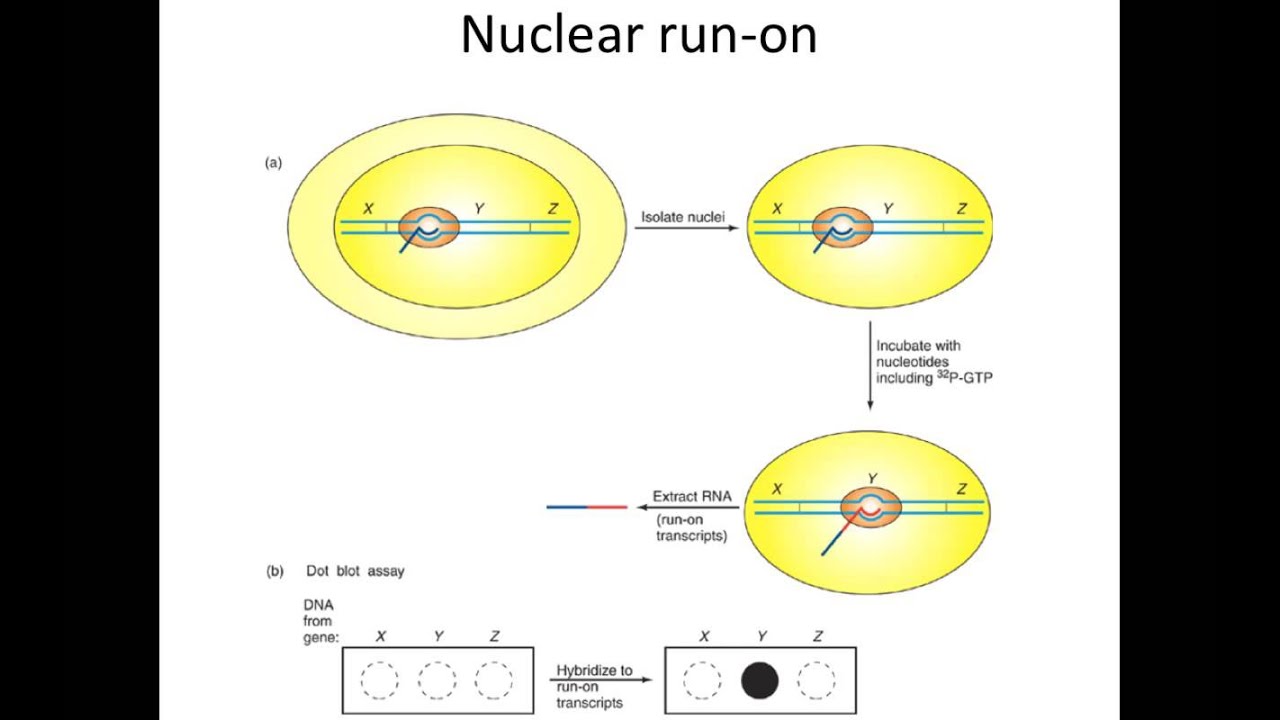

![II] Molecular Techniques for Studying Gene Expression - ppt download II] Molecular Techniques for Studying Gene Expression - ppt download](https://slideplayer.com/slide/6359784/22/images/60/Nuclear+Run-on+Transcription+Assay.jpg)